Christine Farrance , Accugenix 09.11.15
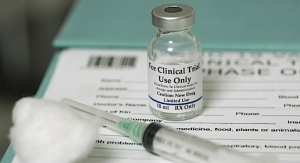
Few of us would ever get a vaccination or unseal a bottle of contact solution if we thought it was contaminated, so responsible manufacturers devote considerable resources making sure bacteria and fungi don’t end up in the supply chain.
While the regulations pertaining to control of microbiological contamination have remained fairly constant over the last decade, the warning letters related to this topic have increased steadily. In fact, whether it’s medical devices, pharmaceuticals or biologics, contamination is among the top areas of risk to public safety.
The headlines are filled with examples of stories of what can happen when pathogens run amok—molds growing in steroid vials and “clean rooms,” bacteria in sterile water filters and superbugs ending up on endoscopes. While it’s a fool’s errand to know every microorganism lurking in our midst—there are tens of thousands, perhaps millions of bacterial species living on Earth—biomedical companies have a moral and legal responsibility to be as proactive as possible about the ones that we are aware of and that spell trouble.
The cornerstone of this effort is environmental monitoring which comprises a biological surveillance system that enables companies to quickly identify organisms that pass through or live in their facilities before they had an opportunity to contaminate the entire “supply chain.” Good environmental monitoring helps avoid the kinds of in-house disasters that lead to costly fines and recalls from regulatory agencies. More importantly, it helps keep the public safe from harm and instills confidence in both the company and regulators alike that the products have been properly tested.
The process is getting faster and more precise thanks to the increased use of genotypic technologies and rapid molecular tests. Companies can now gain a more complete dossier of any number of these wily stealth agents that turn up in their facilities. Companies are also able to spot trends more quickly that spell trouble or, conversely, show their quality control measures are working. Genotypic tools allow companies to drill deeply into an organism’s family tree and identify subtle differences that can explain, sometimes in excruciating detail, how one family member behaves compared to another. The delivery of information is occurring faster as well.
While these are all good and welcome advances, the challenge remains convincing companies which advances in microbial identification are worth the time, effort and money. It’s not always clear which platforms are the best ones to use to manage the risk of tracking microbial identification.
US Food and Drug Administration (FDA) guidelines spelled out here and here, outline the good manufacturing practices (GMP) that companies need to follow to make sure their products are adequately tested and free of objectionable organisms. The manufacturer is responsible for establishing specifications for identity and purity, as well as limits on those types of contaminants that may adulterate the finished product. These guidelines may sound simple and straightforward, but with every step in the manufacturing process at risk for contamination following these regulatory recommendations is actually quite cumbersome.
To modernize the process, the FDA launched Pharmaceutical cGMP in 2003 to encourage the industry to develop and embrace innovative quality control strategies that improve drug manufacturing processes. The initiative includes a 13-year-old guidance document known as Process Analytical Technology (PAT) that joins modern analytical technology with information management and analysis to quickly identify and control conditions that affect the manufacturing process.
For cultural and historical reasons, the pharmaceutical industry has been slow to implement rapid technologies, though it is starting to move toward implementing some of these practices for in-process testing, bioburden assessment, sterility testing, microbial limit testing and other areas.
Tracking and trending
So what do we really mean when we use the term environmental monitoring? On the most basic level, environmental monitoring provides a baseline profile of a manufacturing environment and enables companies to promptly identify sites at risk of contaminating a product. It also acts as an early warning system by alerting facilities when contaminants have exceeded the acceptable exposure limits. The statistical analyses conducted by environmental monitoring programs over time can also flag deviations in and conformance to acceptable limits. The data also highlight trends by comparing the number of colonies and evaluating recurrences of species.
In a lot of ways, environmental monitoring is a company’s home-grown eavesdropping system, a way of identifying ominous threats. Figuring out if the chatter is authentic and then how to dispense with the threat can be a challenge for companies continually searching for ways to improve their bottom line. Some of these methods are more expensive or complex than others.
The identification of microorganisms can be done through different processes, each with its own level of accuracy and reproducibility. Accurate and consistent identification methods are critical in managing risk by yielding data that allow for comprehensive and reliable tracking and trending during routine monitoring of and investigations into excursions. Accuracy of identification and strain typing is key and dependent on the method used to generate and interpret the data as well as the library database used as a reference.
Exposing secret agents
So how do such facilities ID organisms? For decades, Gram staining has been used by laboratories to differentiate bacterial species into two large groups—Gram-negative and Gram-positive—and, although error prone, this diagnostic tool continues to be the first step in identifying organisms in clinical and research settings. A common microbial identification method relies on Gram staining and is an automated identification system designed to assess microwells of thin plastic identification cards about the size of a standard playing card. One must first perform a Gram stain reaction and perform a microscopic examination to determine the cell morphology of the bacteria which determines which card should be used. Each card includes between 30-64 wells that contain different dehydrated biochemical substrates—chemical compounds that trigger enzymatic reactions—which are used as a kind of Geiger counter or Morse code in identifying microorganisms.
But most bacteria cannot be nailed down using Gram-staining and phenotypic methods, and as the saying goes close only counts in horseshoes. So since the mid-1990s, the standard tool used by scientists to identify bacteria has been to sequence the small subunit (16S) ribosomal RNA gene (the 16S rDNA). Sequencing the 16S rDNA allows for comparative analysis of published sequences in microbial databases; you can determine if you have uncovered something that has been seen before or something that looks to be entirely new. In recent years, facilities have capitalized on technological advances and begun using MALDI-TOF MS for routine identifications (for matrix-assisted laser desorption/ionization-time of flight mass spectrometry). This rapid and accurate testing method uses formic acid, matrix and a laser to break apart bacterial specimens and vaporizes the ionized proteins. The protein spectral patterns of the unknown are then compared to a library of known organisms’ spectra for identification.
While 16S rDNA has revolutionized the classification of bacteria, it has also elucidated the misclassification of numerous bacteria that were named before the advent of DNA sequencing. Historically, species would be classified as different based on phenotypic or clinical properties. These properties can vary based on how a bug is grown or what host it has been exposed to, but the DNA stays the same. Once the 16S rDNA sequences were derived, the story got muddled and different species had the same name or the same species had different names. The challenge is to unravel the confusion that is bacterial taxonomy.
So one thing we have done is to develop and maintain multiple identification libraries as references for different kingdoms of organisms (bacteria and fungi) so that our customers—pharmaceutical, medical device, dietary supplement and personal care product manufacturers—know what bugs might be lurking in their facilities or products and how to respond appropriately. We continuously comb the scientific literature for examples of new and reclassified species that might be good candidates to add to our library. We vet each organism to make sure its nomenclature is right. We may order the type strain from one of the international culture collections, sequence the 16S or ITS2 rDNA region and evaluate its phylogeny or taxonomy.
The question is not just academic. The federal government requires that companies monitor their environment and keep track of what is in that environment. If you can’t get an accurate name on an organism, it makes it difficult to be confident in the results obtained from routine microbial surveillance. Following these regulations effectively is a great challenge for manufactures of critical health care products because every aspect of production is at continual risk of being a vector for microbial contamination of the product as depicted below.
And when an industry is allowed to operate in an unregulated environment, as was the case with the New England Compounding Center, the consequences from contamination can be catastrophic.
Fungal tragedy
The highly publicized 2012 FDA inspection of NECC discovered that a quarter of the MPA steroid (methylprednisolone acetate) injections in one bin contained “greenish black foreign matter” and identified several clean rooms that had bacterial or mold growth. Floor mats near sterile drug-mixing areas were “visibly soiled with assorted debris” and a leak from a nearby boiler created an “environment susceptible to contaminant growth.”
The tainted steroid medications sparked an unprecedented fungal meningitis outbreak in 2012, killing 64 people and sickening over 680 others. A black mold, Exserohilum rostratum, was found in 151 cases and 23 additional species of fungi were identified from patients. Also unprecedented was that this fungus wasn’t known to make people sick, let alone cause a deadly inflammatory disease. The outbreak led to the shutdown and bankruptcy of the Framingham-based company, and 14 individuals have since been arrested in a criminal probe spanning several states.
NECC is not the only scary example of microbial contamination in recent years. Specialized endoscopes used to treat disorders of the digestive tract were found to be the cause of widespread infection with the “super bug” carbapenem-resistant Enterobacteriaceae (CRE) that can kill up to 40% of the people it infects.
Proper environmental monitoring should help companies to avoid incidents like these by delivering an accurate snapshot of what organisms could be in or around their product. But environmental monitoring on any level won’t work unless a facility uses accurate and reproducible methods of identification. Yet because the taxonomy of microorganisms is not straightforward, choosing the best identification method is critical. Phylogenetic analyses using DNA sequences, combined with a curated reference library, can elucidate the identification and the taxonomic position of an organism with the most accuracy.
Fungal organisms illustrate how complicated this process can be. Of the 112 recalls by the FDA between 2000 and 2010, contamination by mold and yeast was found in 21% of the samples. But possibly because it’s so hard to obtain an ID using standard tools, molds and yeasts are not generally speciated. Any contamination event in a sterile product is unacceptable, but even with nonsterile products, especially respiratory products, mold will cause serious issues. So the emphasis should be on genotypic methods for ID.
Accurately identifying the organism to the species, and even strain level permits the company to track the potential origin of contamination. It also avoids delays in product release and completion of investigations and permits verification of final product identity. Strain level characterization using genotypic methods is critical in the case of a major excursion or sterility failure and for routine typing of isolates from events exceeding alert and action levels. The data can be used to build a custom library database of sequences and serves as a point of reference that can then be used to monitor the facility.
The data gathered in a well-designed and executed environmental monitoring program will pay off in the end. Reliable microbial identification and strain typing methods are needed to give consistent and accurate results for trending and tracking of organisms. Methods which provide an inconsistent ID or no ID for the same isolate are not useful for tracking isolates to their source or for generating trending reports and can lead to misdirected remediation efforts.
Christine Farrance, PhD, is the director of research and development for the Accugenix portfolio of services at Charles River Laboratories. Accugenix is based in Newark, Delaware.
While the regulations pertaining to control of microbiological contamination have remained fairly constant over the last decade, the warning letters related to this topic have increased steadily. In fact, whether it’s medical devices, pharmaceuticals or biologics, contamination is among the top areas of risk to public safety.
The headlines are filled with examples of stories of what can happen when pathogens run amok—molds growing in steroid vials and “clean rooms,” bacteria in sterile water filters and superbugs ending up on endoscopes. While it’s a fool’s errand to know every microorganism lurking in our midst—there are tens of thousands, perhaps millions of bacterial species living on Earth—biomedical companies have a moral and legal responsibility to be as proactive as possible about the ones that we are aware of and that spell trouble.
The cornerstone of this effort is environmental monitoring which comprises a biological surveillance system that enables companies to quickly identify organisms that pass through or live in their facilities before they had an opportunity to contaminate the entire “supply chain.” Good environmental monitoring helps avoid the kinds of in-house disasters that lead to costly fines and recalls from regulatory agencies. More importantly, it helps keep the public safe from harm and instills confidence in both the company and regulators alike that the products have been properly tested.
The process is getting faster and more precise thanks to the increased use of genotypic technologies and rapid molecular tests. Companies can now gain a more complete dossier of any number of these wily stealth agents that turn up in their facilities. Companies are also able to spot trends more quickly that spell trouble or, conversely, show their quality control measures are working. Genotypic tools allow companies to drill deeply into an organism’s family tree and identify subtle differences that can explain, sometimes in excruciating detail, how one family member behaves compared to another. The delivery of information is occurring faster as well.
While these are all good and welcome advances, the challenge remains convincing companies which advances in microbial identification are worth the time, effort and money. It’s not always clear which platforms are the best ones to use to manage the risk of tracking microbial identification.
US Food and Drug Administration (FDA) guidelines spelled out here and here, outline the good manufacturing practices (GMP) that companies need to follow to make sure their products are adequately tested and free of objectionable organisms. The manufacturer is responsible for establishing specifications for identity and purity, as well as limits on those types of contaminants that may adulterate the finished product. These guidelines may sound simple and straightforward, but with every step in the manufacturing process at risk for contamination following these regulatory recommendations is actually quite cumbersome.
To modernize the process, the FDA launched Pharmaceutical cGMP in 2003 to encourage the industry to develop and embrace innovative quality control strategies that improve drug manufacturing processes. The initiative includes a 13-year-old guidance document known as Process Analytical Technology (PAT) that joins modern analytical technology with information management and analysis to quickly identify and control conditions that affect the manufacturing process.
For cultural and historical reasons, the pharmaceutical industry has been slow to implement rapid technologies, though it is starting to move toward implementing some of these practices for in-process testing, bioburden assessment, sterility testing, microbial limit testing and other areas.
Tracking and trending
So what do we really mean when we use the term environmental monitoring? On the most basic level, environmental monitoring provides a baseline profile of a manufacturing environment and enables companies to promptly identify sites at risk of contaminating a product. It also acts as an early warning system by alerting facilities when contaminants have exceeded the acceptable exposure limits. The statistical analyses conducted by environmental monitoring programs over time can also flag deviations in and conformance to acceptable limits. The data also highlight trends by comparing the number of colonies and evaluating recurrences of species.
In a lot of ways, environmental monitoring is a company’s home-grown eavesdropping system, a way of identifying ominous threats. Figuring out if the chatter is authentic and then how to dispense with the threat can be a challenge for companies continually searching for ways to improve their bottom line. Some of these methods are more expensive or complex than others.
The identification of microorganisms can be done through different processes, each with its own level of accuracy and reproducibility. Accurate and consistent identification methods are critical in managing risk by yielding data that allow for comprehensive and reliable tracking and trending during routine monitoring of and investigations into excursions. Accuracy of identification and strain typing is key and dependent on the method used to generate and interpret the data as well as the library database used as a reference.
Exposing secret agents
So how do such facilities ID organisms? For decades, Gram staining has been used by laboratories to differentiate bacterial species into two large groups—Gram-negative and Gram-positive—and, although error prone, this diagnostic tool continues to be the first step in identifying organisms in clinical and research settings. A common microbial identification method relies on Gram staining and is an automated identification system designed to assess microwells of thin plastic identification cards about the size of a standard playing card. One must first perform a Gram stain reaction and perform a microscopic examination to determine the cell morphology of the bacteria which determines which card should be used. Each card includes between 30-64 wells that contain different dehydrated biochemical substrates—chemical compounds that trigger enzymatic reactions—which are used as a kind of Geiger counter or Morse code in identifying microorganisms.
But most bacteria cannot be nailed down using Gram-staining and phenotypic methods, and as the saying goes close only counts in horseshoes. So since the mid-1990s, the standard tool used by scientists to identify bacteria has been to sequence the small subunit (16S) ribosomal RNA gene (the 16S rDNA). Sequencing the 16S rDNA allows for comparative analysis of published sequences in microbial databases; you can determine if you have uncovered something that has been seen before or something that looks to be entirely new. In recent years, facilities have capitalized on technological advances and begun using MALDI-TOF MS for routine identifications (for matrix-assisted laser desorption/ionization-time of flight mass spectrometry). This rapid and accurate testing method uses formic acid, matrix and a laser to break apart bacterial specimens and vaporizes the ionized proteins. The protein spectral patterns of the unknown are then compared to a library of known organisms’ spectra for identification.
While 16S rDNA has revolutionized the classification of bacteria, it has also elucidated the misclassification of numerous bacteria that were named before the advent of DNA sequencing. Historically, species would be classified as different based on phenotypic or clinical properties. These properties can vary based on how a bug is grown or what host it has been exposed to, but the DNA stays the same. Once the 16S rDNA sequences were derived, the story got muddled and different species had the same name or the same species had different names. The challenge is to unravel the confusion that is bacterial taxonomy.
So one thing we have done is to develop and maintain multiple identification libraries as references for different kingdoms of organisms (bacteria and fungi) so that our customers—pharmaceutical, medical device, dietary supplement and personal care product manufacturers—know what bugs might be lurking in their facilities or products and how to respond appropriately. We continuously comb the scientific literature for examples of new and reclassified species that might be good candidates to add to our library. We vet each organism to make sure its nomenclature is right. We may order the type strain from one of the international culture collections, sequence the 16S or ITS2 rDNA region and evaluate its phylogeny or taxonomy.
The question is not just academic. The federal government requires that companies monitor their environment and keep track of what is in that environment. If you can’t get an accurate name on an organism, it makes it difficult to be confident in the results obtained from routine microbial surveillance. Following these regulations effectively is a great challenge for manufactures of critical health care products because every aspect of production is at continual risk of being a vector for microbial contamination of the product as depicted below.
And when an industry is allowed to operate in an unregulated environment, as was the case with the New England Compounding Center, the consequences from contamination can be catastrophic.
Fungal tragedy
The highly publicized 2012 FDA inspection of NECC discovered that a quarter of the MPA steroid (methylprednisolone acetate) injections in one bin contained “greenish black foreign matter” and identified several clean rooms that had bacterial or mold growth. Floor mats near sterile drug-mixing areas were “visibly soiled with assorted debris” and a leak from a nearby boiler created an “environment susceptible to contaminant growth.”
The tainted steroid medications sparked an unprecedented fungal meningitis outbreak in 2012, killing 64 people and sickening over 680 others. A black mold, Exserohilum rostratum, was found in 151 cases and 23 additional species of fungi were identified from patients. Also unprecedented was that this fungus wasn’t known to make people sick, let alone cause a deadly inflammatory disease. The outbreak led to the shutdown and bankruptcy of the Framingham-based company, and 14 individuals have since been arrested in a criminal probe spanning several states.
NECC is not the only scary example of microbial contamination in recent years. Specialized endoscopes used to treat disorders of the digestive tract were found to be the cause of widespread infection with the “super bug” carbapenem-resistant Enterobacteriaceae (CRE) that can kill up to 40% of the people it infects.
Proper environmental monitoring should help companies to avoid incidents like these by delivering an accurate snapshot of what organisms could be in or around their product. But environmental monitoring on any level won’t work unless a facility uses accurate and reproducible methods of identification. Yet because the taxonomy of microorganisms is not straightforward, choosing the best identification method is critical. Phylogenetic analyses using DNA sequences, combined with a curated reference library, can elucidate the identification and the taxonomic position of an organism with the most accuracy.
Fungal organisms illustrate how complicated this process can be. Of the 112 recalls by the FDA between 2000 and 2010, contamination by mold and yeast was found in 21% of the samples. But possibly because it’s so hard to obtain an ID using standard tools, molds and yeasts are not generally speciated. Any contamination event in a sterile product is unacceptable, but even with nonsterile products, especially respiratory products, mold will cause serious issues. So the emphasis should be on genotypic methods for ID.
Accurately identifying the organism to the species, and even strain level permits the company to track the potential origin of contamination. It also avoids delays in product release and completion of investigations and permits verification of final product identity. Strain level characterization using genotypic methods is critical in the case of a major excursion or sterility failure and for routine typing of isolates from events exceeding alert and action levels. The data can be used to build a custom library database of sequences and serves as a point of reference that can then be used to monitor the facility.
The data gathered in a well-designed and executed environmental monitoring program will pay off in the end. Reliable microbial identification and strain typing methods are needed to give consistent and accurate results for trending and tracking of organisms. Methods which provide an inconsistent ID or no ID for the same isolate are not useful for tracking isolates to their source or for generating trending reports and can lead to misdirected remediation efforts.
Christine Farrance, PhD, is the director of research and development for the Accugenix portfolio of services at Charles River Laboratories. Accugenix is based in Newark, Delaware.